Introduction
Advancements in RNA therapeutics are increasing the demand for high-quality synthetic oligonucleotides including siRNA (small interfering RNA) and sgRNA (single guide RNA). While conventional solid-phase oligonucleotide synthesis (SPOS) remains the primary method for siRNA and sgRNA production, its limitations in scalability, sustainability, quality, and cost hinder its ability to meet the anticipated market demands. Chemoenzymatic ligation, a hybrid approach integrating chemical synthesis with enzymatic ligation, is emerging as a game changing solution to these challenges. This article explores the limitations of SPOS, and provides an overview of chemoenzymatic ligation, its mechanism, and comparative advantages in siRNA and sgRNA synthesis.
Limitations and Challenges of the SPOS Process for siRNA and sgRNA Manufacture
Driven primarily by anticipated approvals in the cardiovascular disease space, it is estimated that global demand for siRNA API may total tens of tons a decade from now. There are significant challenges faced by conventional SPOS1 (we refer to this as Generation 1 technology), in scaling to meet this volume. Current SPOS processes use flow through synthesizers which are limited to batch sizes of ~10 kg (manufacturing larger volumes requires scaling out), consume large volumes of reagents that generate substantial chemical waste, and are costly. sgRNA molecules, which are important components of gene editing drugs, are typically ≥100 nucleotides long. Using SPOS, sgRNA is synthesized by successive addition of nucleotide subunits using the phosphoramidite method, which presents many opportunities for impurities to form and accumulate. These impurities are challenging to control during synthesis and to remove during purification, leading to low purity, potential issues with batch-to-batch consistency, and very high cost.
Chemoenzymatic Ligation: An Overview
Chemoenzymatic ligation (we refer to this as Generation 2 technology), uses a combination of chemical and enzymatic approaches to synthesize oligonucleotides. This method utilizes oligonucleotide fragments (also commonly called blockmers), which are synthesized by SPOS and joined together with an enzyme called ligase to produce full-length RNA molecules. There are three key mechanistic steps in the enzymatic ligation reaction2, as shown in figure 1a, which result in some important properties. Firstly, some impurities are rejected during the process because they’re not viable substrates for the ligation reaction. Secondly, ligases generally act with high accuracy, and yield and rate of reaction can be optimized by tweaking the various process parameters. Lastly, the reaction is stereospecific and a phosphorothioate diester with Rp stereochemistry is generated when a fragment with pro-chiral 5’-thiophosphate is used as a substrate3. This should be carefully considered if ligation is to be applied for antisense oligonucleotide (ASO) synthesis.
siRNA Synthesis Using Sticky End Ligation
Pioneered by Khorana et al4, so-called sticky end ligation is a key step in DNA cloning techniques and there have been reports on its application in siRNA synthesis5,6. The sticky end ligation process used by Hongene for synthesis of siRNA is shown in figure 1b. There are several advantages of sticky end ligation compared with conventional SPOS:
- Quality: The selective rejection of shortmer impurities (oligonucleotides lacking one or more nucleotides) during ligation enhances final product purity and batch-to-batch consistency compared to conventional SPOS. Additionally, different strategies can be employed depending on customer purity and cost expectations. For example, oligonucleotide fragments can be purified by chromatography (higher quality and cost) or simply desalted (lower purity and cost) prior to ligation.
- Yield, cost, and environmental sustainability: Modular ligation of oligonucleotide fragments is performed in near quantitative yields from oligonucleotide fragments which are synthesized in high yield using conventional SPOS. The overall process is higher yielding, leading to reduced cost and improved sustainability.
- Scalability: The ligation reaction is performed in batch reactors that can be disposable, providing a more scalable solution for GMP manufacturing.
sgRNA Synthesis Using Splinted Ligation
Long RNA molecules can be synthesized by splinted ligation7, and its application for synthesis of chemically modified RNA up to 120 nucleotides long has been recently reported8. At Hongene we have developed a process based on splinted ligation for assembling sgRNA and other long RNA constructs as shown in figure 1c. The advantages described for sticky end ligation also apply for splinted ligation. In addition, by virtue of its modularity, splinted ligation can be applied to synthesis of long chemically modified RNA molecules such as those required for prime editing and related technologies, without compromising quality or yield (see Figure 1 below).
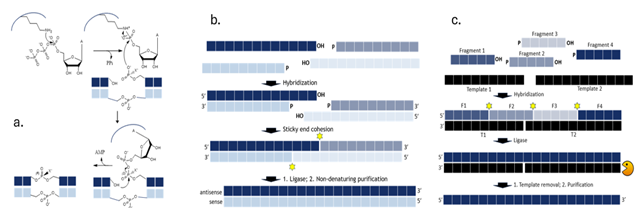
Figure 1. a) The enzymatic ligation reaction involves 3 mechanistic steps: 1) Adenylylation: The enzyme’s lysine residue reacts with ATP provided in the reaction mix to form an adenylylated intermediate, releasing pyrophosphate, 2) Phosphate transfer: The adenylylated intermediate reacts with the 5′-phosphate of one fragment, creating a bridged diphosphate, 3) Ligation: The adjoining oligonucleotide fragment’s 3′-OH group reacts with the bridged diphosphate, releasing AMP and sealing the junction; b) Hongene’s sticky end ligation process involves hybridization of four partially overlapping and complementary oligonucleotide fragments to form two duplexes with complementary single stranded overhangs that also hybridize. The two resulting junctions are joined by RNA ligase to generate hybridized duplex siRNA that can be readily purified by non-denaturing chromatography; c) In Hongene’s splinted ligation process DNA splints hybridize with overlapping complementary RNA fragments, facilitating precise joining of the resulting junctions with RNA ligase. The crude post-ligation reaction mixture is then treated with DNase I to digest the splints followed by chromatographic purification to furnish sgRNA with high yield and purity.
Status of siRNA and sgRNA Manufacturing at Hongene Using Chemoenzymatic Ligation Technology
Our experience to date indicates that siRNA chemistries typically found in clinical molecules (including ligands such as GalNAc) are well tolerated during ligation. We are actively conducting proof of concept work with several collaborators and have completed our first GMP manufacturing of a development candidate for clinical studies (table 1).
Table 1. Examples of oligonucleotide products synthesized by chemoenzymatic ligation at Hongene
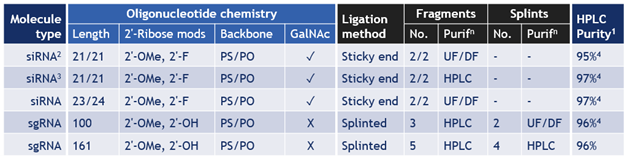
1) Denaturing UPLC method; 2) non-GMP engineering batch; 3) GMP batch same siRNA as 1st row; and 4) Data verified by customer.
Future Perspectives
The transition to chemoenzymatic synthesis marks a technological step change in oligonucleotide manufacturing. While current applications demonstrate significant promise, further innovations will maximize its potential. Advances in enzyme engineering, process optimization, and integration with improved SPOS and liquid-phase technologies9 for more efficient synthesis of oligonucleotide fragments will shape the future of RNA therapeutics production.
In parallel, development of fully enzymatic synthesis methods10–13 (we refer to this as Generation 3 technology), hold the promise of bypassing chemical synthesis entirely, offering a sustainable and efficient alternative for oligonucleotide production.
Conclusion
Chemoenzymatic ligation is poised to change the landscape of oligonucleotide manufacturing. Its ability to deliver, high purity siRNA and sgRNA with reduced cost, environmental impact, and scalable production processes positions it as a viable technology for meeting the anticipated future demand for RNA drugs.
About the Author

![]() David Butler is the Chief Technology Officer at Hongene Biotech Corporation.
David Butler is the Chief Technology Officer at Hongene Biotech Corporation.
Find him on LinkedIn.
References
(1) Sanghvi, Y. S.; Ferrazzano, L.; Cabri, W.; Tolomelli, A. Sustainable Approaches in Solid-Phase Oligonucleotide Synthesis: Current Status and Future Directions. In Sustainability in Tides Chemistry; Tolomelli, A., Ferrazzano, L., Cabri, W., Eds.; Royal Society of Chemistry, 2024; pp 228–247. https://doi.org/10.1039/9781837674541-00228.
(2) Nandakumar, J.; Shuman, S.; Lima, C. D. RNA Ligase Structures Reveal the Basis for RNA Specificity and Conformational Changes That Drive Ligation Forward. Cell 2006, 127 (1), 71–84. https://doi.org/10.1016/j.cell.2006.08.038.
(3) Bryant, F. R.; Benkovic, S. J. Phosphorothioate Substrates for T4 RNA Ligase. Biochemistry 1982, 21 (23), 5877–5885. https://doi.org/10.1021/bi00266a023.
(4) Khorana, H. G. Nucleic Acid Synthesis. Pure Appl. Chem. 1968, 17 (3), 349–382. https://doi.org/10.1351/pac196817030349.
(5) Paul, S.; Gray, D.; Caswell, J.; Brooks, J.; Ye, W.; Moody, T. S.; Radinov, R.; Nechev, L. Convergent Biocatalytic Mediated Synthesis of siRNA. ACS Chem. Biol. 2023. https://doi.org/10.1021/acschembio.3c00071.
(6) Kajimoto, S.; Ohashi, M.; Hagiwara, Y.; Takahashi, D.; Mihara, Y.; Motoyama, T.; Ito, S.; Nakano, S. Enzymatic Conjugation of Modified RNA Fragments by Ancestral RNA Ligase AncT4_2. Appl. Environ. Microbiol.88 (23), e01679-22. https://doi.org/10.1128/aem.01679-22.
(7) Moore, M. J.; Query, C. C. [7] Joining of RNAs by Splinted Ligation. In Methods in Enzymology; RNA – Ligand Interactions, Part A; Academic Press, 2000; Vol. 317, pp 109–123. https://doi.org/10.1016/S0076-6879(00)17009-0.
(8) Sabat, N.; Stämpfli, A.; Hanlon, S.; Bisagni, S.; Sladojevich, F.; Püntener, K.; Hollenstein, M. Template-Dependent DNA Ligation for the Synthesis of Modified Oligonucleotides. Nat. Commun. 2024, 15 (1), 8009. https://doi.org/10.1038/s41467-024-52141-8.
(9) Ferrazzano, L.; Rapisarda, L. Sustainable Approaches in Liquid-Phase Oligonucleotide Synthesis (LPOS). In Sustainability in Tides Chemistry; Tolomelli, A., Ferrazzano, L., Cabri, W., Eds.; Royal Society of Chemistry, 2024; pp 248–272. https://doi.org/10.1039/9781837674541-00248.
(10) Moody, E. R.; Obexer, R.; Nickl, F.; Spiess, R.; Lovelock, S. L. An Enzyme Cascade Enables Production of Therapeutic Oligonucleotides in a Single Operation. Science 2023, 380 (6650), 1150–1154. https://doi.org/10.1126/science.add5892.
(11) Sabat, N.; Katkevica, D.; Pajuste, K.; Flamme, M.; Stämpfli, A.; Katkevics, M.; Hanlon, S.; Bisagni, S.; Püntener, K.; Sladojevich, F.; Hollenstein, M. Towards the Controlled Enzymatic Synthesis of LNA Containing Oligonucleotides. Front. Chem. 2023, 11, 1161462. https://doi.org/10.3389/fchem.2023.1161462.
(12) Wiegand, D. J.; Rittichier, J.; Meyer, E.; Lee, H.; Conway, N. J.; Ahlstedt, D.; Yurtsever, Z.; Rainone, D.; Kuru, E.; Church, G. M. Template-Independent Enzymatic Synthesis of RNA Oligonucleotides. Nat. Biotechnol. 2024, 1–11. https://doi.org/10.1038/s41587-024-02244-w.
(13) Forget, S.; Krawczyk, M.; Knight, A.; Ching, C.; Copeland, R.; Mahmoodi, N.; Mayo, M.; Nguyen, J.; Tan, A.; Miller, M.; Vroom, J.; Lutz, S. Evolving a High-Performance Terminal Deoxynucleotidyl Transferase for Enzymatic DNA Synthesis. bioRxiv June 27, 2024, p 2024.06.24.600205. https://doi.org/10.1101/2024.06.24.600205.